
Evaluating BERT to build a scientific sentiment model of sentences
“We have no evidence of…” can mean different things to different people. For example, “we have no evidence of transmission of SARS-Cov-2 between humans and cats” might have the intended meaning of “we don’t know yet if there is or is not transmission.” If, however, especially among members of the general public who don’t usually read scientific literature, it could readily be interpreted as “transmission doesn’t happen.”
This is not just another “cats on the internet” meme. Many people have been using search engines to help navigate life during the coronavirus pandemic. General search engine results in March 2020 might have led a user to believe that it was completely safe to let strangers interact with their pets during quarantine, although at least one paper had been published demonstrating ACE-2 receptor expression in ferrets and cats to be a possible COVID19 transmission risk, in advance of additional evidence and ultimately the CDC’s confirmation that humans can pass this disease to their feline friends.
The interplay between academic research, publications, and search engines is rarely simple. For example, it can be hard to interpret what it means when there are no relevant results. Is there truly no information on this topic? Or should you be asking a different question? Even a casual search engine user knows that small changes in a search query can dramatically alter results. For instance, there are many different ways to search for the fatality rate for COVID-19, including case fatality ratio (CFR), infection fatality ratio (IFR), or death rate. Each returns a different result. In fact, a query just on IFRs will yield different results based on the population, frequency of testing, and potential over or under-reporting of deaths from COVID-19. It remains an open question how best to summarize such varying, conflicting results, and ensure that users are getting answers to the question they meant to ask, rather than what they actually queried.

In our previous post, we discussed some of the challenges with using and evaluating knowledge discovery tools, such as Question-and-Answer systems, especially in the context of COVID-19. While we were able to compare submissions to a recent Kaggle competition against a minimal baseline we created, we still don’t know if any of these submissions are correct, complete, or ultimately utile for researchers and policymakers. We couldn’t find any submission besides our own that made an effort to evaluate the quality of the results it returned.
These kinds of COVID-19 specific search engines and knowledge discovery tools produced by research groups and startups suffer from what the software testing community calls the Oracle Problem: you cannot evaluate a system in which there are no known, correct outputs for a range of inputs. COVID-19 is a new disease, where today’s fact may become tomorrow’s fallacy, and yet we needed answers yesterday. How do we begin to evaluate knowledge discovery tools for correctness, completeness, and utility in this domain? Our future post in this series will discuss our own user study conducted on evaluating knowledge discovery tools, including the ethical implications of relying on crowd-sourcing for this type of problem. Meanwhile, in this post, we’ll continue investigating how we can evaluate these tools in the biomedical domain in the absence of good test oracles.
We’ll expand upon the evaluation of our submission to the Kaggle challenge mentioned above. As we explained in a previous post, our submission performed well against a baseline for finding important, known treatments for COVID19. We built this tool using a novel model of human-centric “efficacy valence” or “scientific sentiment,” a concept that we explain and evaluate below. This post will also discuss the difficulty of evaluating models beyond a minimal baseline, motivating our forthcoming user study as the final post on this topic.
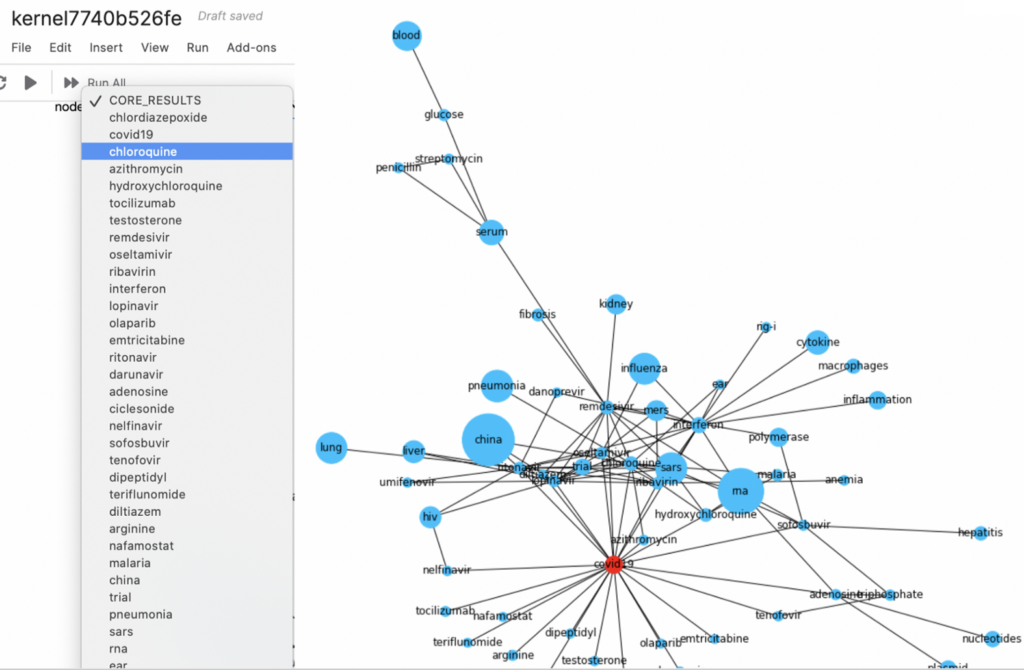
Motivating our scientific sentiment approach to weighting nodes and edges
The CORD-19 Kaggle challenge presented analysts with tens of thousands of academic articles and challenged participants to address a series of COVID19-related questions, including “What do we know about therapeutics?” Participants had to distill the information in these articles into something useful and manageable for humans. Our team decided that we would try to filter sentences in the academic articles that not only mentioned a drug or treatment but also had a higher likelihood of referring to meaningful experimental results and learnings. We dub this weighting scientific sentiment. Other similar works detect causal sentences, as well as explicit and implicit claims, versus correlations and comparisons in biomedical literature.
We hypothesized that rating sentences by their scientific sentiment could yield better performance for our drug discovery tool, at least compared to naïve approaches. In particular, we wanted to try to mimic how a human would judge a sentence for this task. Our approach towards predicting scientific sentiment is similar to work that combines polarity and strength ratings for food-gene-disease networks. Some research goes even deeper, predicting dimensions of focus (scientific, generics, or methodology), evidence (low to high), polarity, certainty (low to high), and directionality for biomedical sentences (predicting five dimensions seems to be popular). These approaches are similar to hedge negation and removal (for example, by user-defined keywords and patterns) and fine-grained factuality analysis of biomedical events.
Although there is a lot of related work in detecting context in biomedical sentences, many of these are rule or pattern-based and for specific types of relations. Instead, we chose to build a sentence rating model based off BERT, which is a revolutionary NLP tool that is able to convert sentences into word embeddings that can be used for downstream machine learning tasks, such as classification. For example, BERT has been used to train models that can predict the sentiment of movie reviews. We chose BERT because of its potential ability to learn context and generalize to efficacy modeling; our goal was to train BERT to take a sentence from an academic article and assign a scientific sentiment ranking. We could then build our knowledge graph using only the highest-ranked sentences.
We should mention that we did not want our analysis to only include strong claims about the relationships between COVID19 and therapeutics. Strong claims may omit drugs that are worth investigating given that COVID19 is a new disease. Relying only on strong claims could also limit our tool’s ability to recommend new drugs for COVID19 because there are currently no known drugs that work for COVID19 (besides perhaps remdesivir at the time this post was written, and dexamethasone shortly thereafter). Instead, we wanted a tool that could mimic how a human might judge information found in articles on COVID19 and related diseases.
Building a scientific sentiment model with BERT
For example, consider the following four anonymized (biomedical keywords replaced with VOID) sentences from various academic papers:
VOIDis similar toVOIDin its action and indications for use and has a lowerVOIDof side effects.- The results showed that
VOIDcould up-regulateVOIDexpression time- and dose- dependently inVOID. VOIDhas been persistent in theVOIDVOIDsince 2012.- Further research on the mechanism of this locus will ultimately provide novel insight into
VOIDmechanisms in thisVOID.
Even with the biomedical keywords that we voided, we hypothesized that the first sentence could be more useful for this type of therapeutic knowledge discovery task than the second, which is almost certainly more useful than the third, and so on.
To test this theory, we built a model of scientific sentiment using a basic BERT model pre-trained on a lowercase vocabulary. We chose this vanilla BERT model to transfer learn from, rather than BioBERT or SciBERT, because we wanted to avoid a model optimized for biomedical Named Entity Recognition and Relation Extraction, since we voided out the biologically-relevant information. Future work could contrast our basic model against one built with BioBERT to see if the biomed-specific Q&A in the latter would increase performance. We also chose not to further pre-train BERT with SQuAD, because we weren’t optimizing for Question-and-Answer, though the latter might also be a project worth further investigation.
With the pre-trained vanilla BERT model at hand, we fine-tuned it on 900 labelled COVID19 article sentences, which we manually annotated for scientific sentiment on a binary scale. For example, in the four example sentences above, the first two were labelled “relevant” and the last two “not relevant.” Before labelling, raw sentences were tagged for Medical Subject Heading (MeSH) keywords and drugs, which were then VOID-ed, so that the model would not accidentally learn to associate scientific sentiment with any particular mention of a disease, treatment, or other biomedical entity. During the human labelling, we also manually recorded any interesting words, such as “found,” “results,” or “significant” that we thought could also be used as a comparison to our BERT-based model for weighting sentences. For example, the presence or absence of these keywords could also hint at the utility of a sentence in an article. After training on nine hundred sentences, our model showed 84% accuracy and an f1-score of 0.87 on the remaining holdout test set of one hundred sentences.
Performance of our BERT-based scientific sentiment knowledge graph
Given that we obtained reasonable performance, even with a limited training dataset, we then used our scientific sentiment model to prune sentences for inclusion in a weighted knowledge graph of COVID19 therapeutics and other core topics. We hypothesized that using scientific sentiment to weight sentences would generate more useful knowledge graphs than more naïve sentence weighting approaches. Using our goal graph from our first blog post, which was generated from a literature summary, we then measured how knowledge graphs generated by these different approaches fared at identifying the baseline therapies in our goal graph.
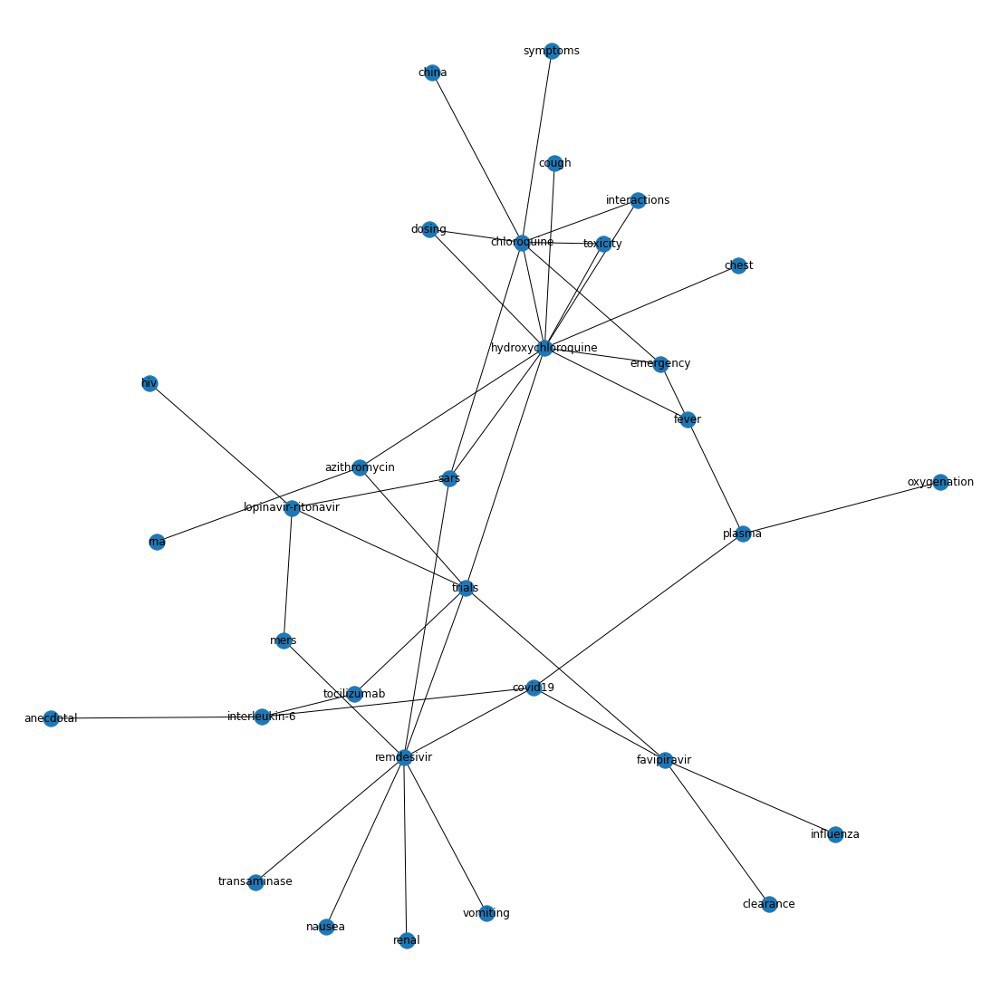
We compared knowledge graphs generated with our scientific sentiment sentence weighting to graphs created with three separate approaches:
- Sentences weighted by how often the drugs they mentioned appeared in our corpus overall;
- Sentences weighted by the citation count of their source articles; and
- Sentences weighted by whether or not they contained human-judged interesting words such as “showed” and “significant.”
We tuned each resulting knowledge graph to contain approximately fifty core nodes and compared the precision of these different weighting schemes at successfully identifying core therapies in our baseline goal graph. We only used sentences from article abstracts, the result of analysis from our previous blog post. Our results are below:
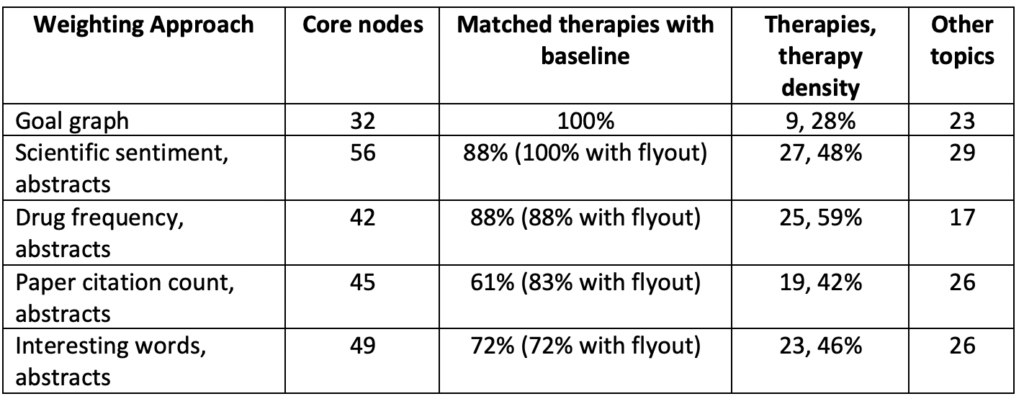
The scientific sentiment weighting was superior to the other sentence weighting schemes at flagging therapies in our goal graph. This is not surprising because BERT better captures context in sentences than a rule-based approach, especially a naïve one that we generated manually. Our scientific sentiment and drug frequency weighting schemes identified the same drugs in their code nodes, but the scientific sentiment model also generated around fifteen more nodes of other related concepts that weren’t drugs (and were not in the goal graph either):
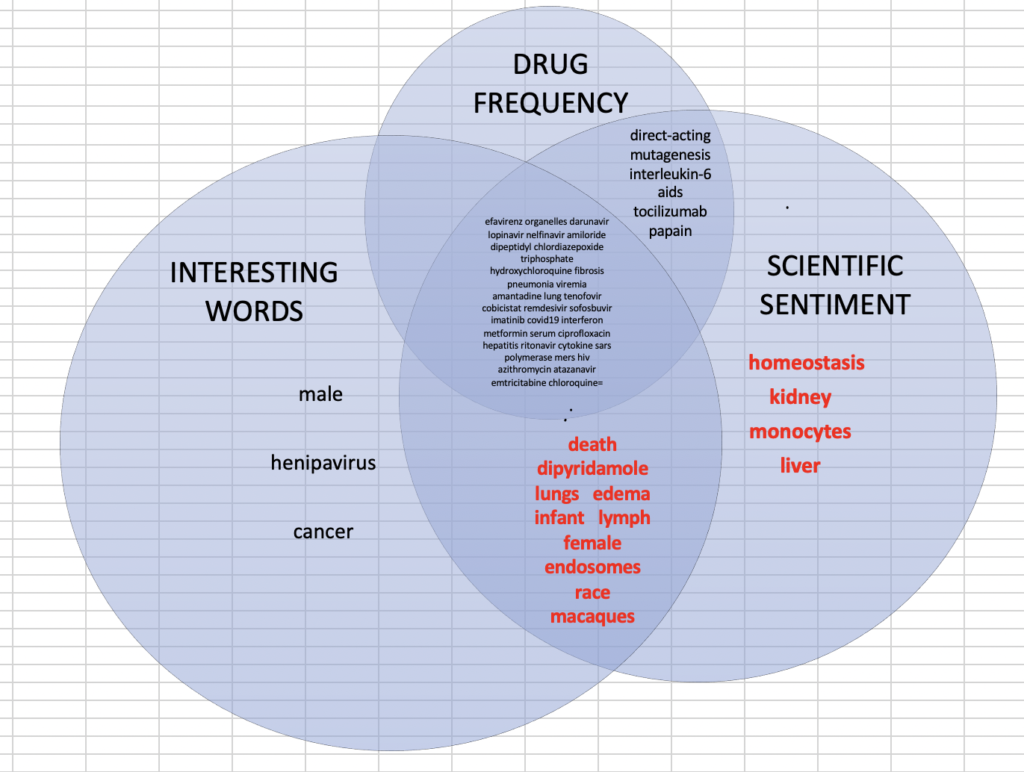
Are these additional, non-drug topics useful to researchers? We think so. The word “female” suggests the different mortality rates we see in women versus men, perhaps because of known sex-based immunity differences and/or other factors. The word “race” may point to the observed differences in mortality for different races, suggesting researchers focus on targeted treatment options and/or policy changes. However, the properties of a correct and useful knowledge discovery tool remain to be explored.
In the meantime, search and knowledge discovery tools continue to emerge at lightning speed, but we still don’t have a way to evaluate their results and methodologies. Although there is now a publicly-available dataset of 124 questions and answers for COVID19, these were mined after the Kaggle competition, using notebooks from two data scientists, with answers that are drawn directly from articles themselves. While these answers were vetted by two expert curators, this method suffers from a key defect: it doesn’t solve the problem of objectively evaluating these notebook submissions or future tools. So can human judgment aid knowledge discovery algorithms? The scientific sentiment score, created by us (humans, after all), proves that it is possible. But we humans will need to devise better means of testing and evaluating these knowledge discovery tools if we’re ever to make progress and not simply more tools.
Towards that end, in our next post in this series, we will discuss the results of a user study we conducted on knowledge discovery tools.

